Project Summary
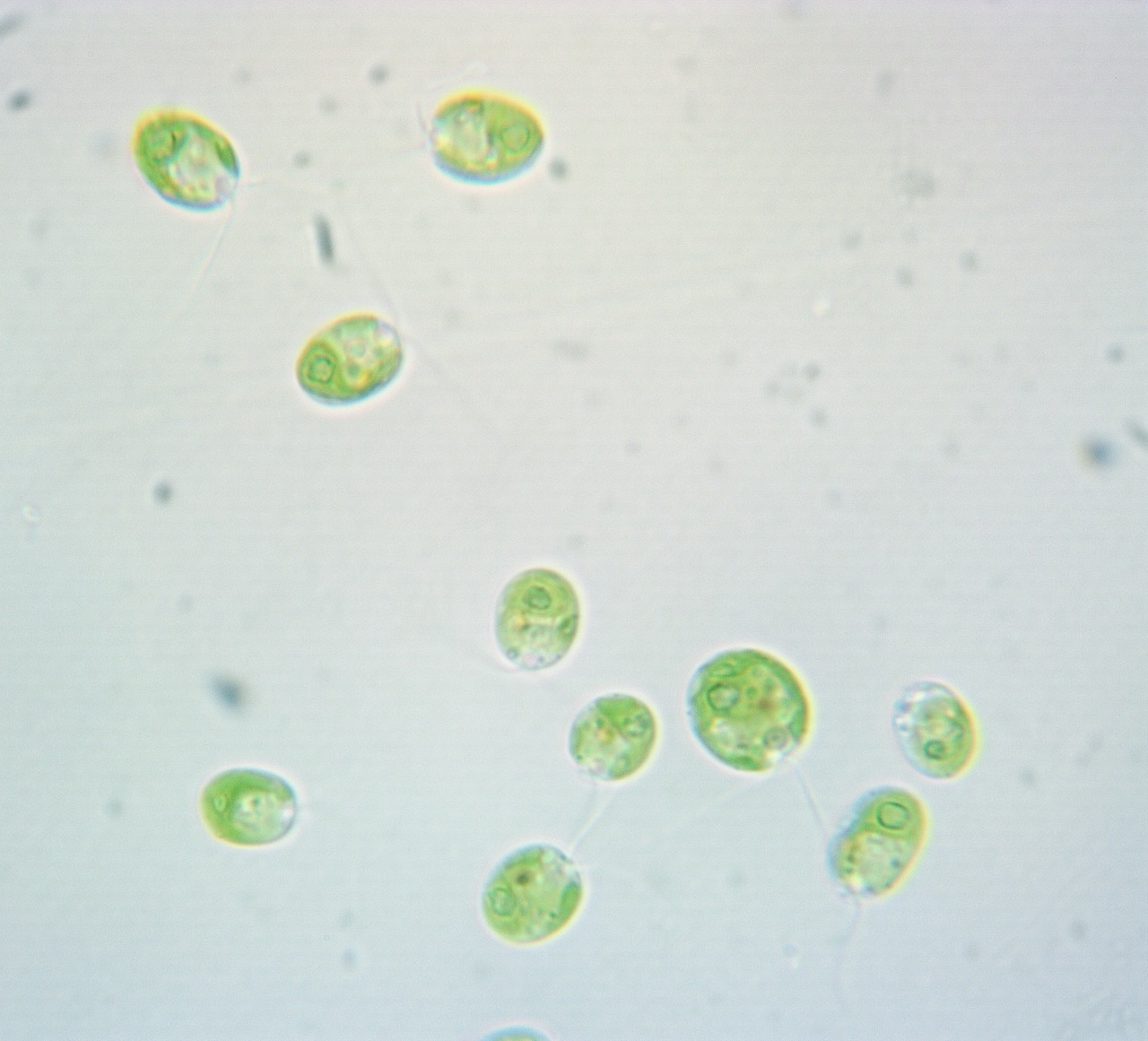
Quorum sensing (QS) is a cell-to-cell communication system that allows unicellular organisms to monitor their population density and synchronize behaviors using signaling molecules. Once a concentration threshold has been reached, the cells collectively alter/express gene expression, leading to a coordinated phenotypic switch, such as biofilm formation and virulence factor production. While predominantly found in prokaryotes, studies have confirmed this phenomenon is conserved among archaea and unicellular eukaryotes. This includes the model photosynthetic unicellular eukaryote Chlamydomonas reinhardtii, in which QS regulates swimming speed. This QS response shows some variation between ecological isolates of this species (strains), suggesting there may be differential responses between lines, as has been observed in some bacterial strains. This information indicates that Chlamydomonas reinhardtii could potentially utilize QS as a broader regulatory mechanism, coordinating changes in motility and metabolism in response to nutrient availability and population density. In this way, QS can facilitate cell distribution and adaptation towards environmental stress factors. With Chlamydomonas reinhardtii, they use photosynthetic pigments such as chlorophylls a and b and carotenoids to capture light for ATP and protect against oxidative stress, QS could help regulate their production as part of an adaptive response, while linking cell density and nutrient conditions to both motility and photosynthetic efficiency, which can lead to cultivation of optimized populations to survival under fluctuating environmental stress. Here, I will leverage an existing microplate assay to determine if there is a correlation between chlorophyll and carotenoid concentration with cell density, supporting a link to quorum sensing. Furthermore, an additional screen phenotypic screening will be conducted, in that model Chlamydomonas reinhardtii will be grown in different media with varying amounts of nitrogen then measured for cell density as well as chlorophyll and carotenoid content. These findings will assist in furthering our understanding of quorum sensing, showing in particular what traits it may influence with Chlamydomonas reinhardtii, specifically if it is with the production of chlorophyll and carotenoids, and if its trigger can be tied to specific environmental factors, such as low or high nutrient availability.
Project Objective
The goal of this study is to evaluate whether QS in Chlamydomonas reinhardtii is regulated by nutrient availability. From testing this, we can either support or refute this claim, thereby uncovering potential applications in biotechnology—such as optimizing algal production for biofuel or pigment synthesis—while providing deeper insight into microbial communication and survival strategies in nutrient-limited environments.
Manufacturing Design Methods
Six cultures were prepared in total, with three controls and three experimental replicates. All six cultures were grown in sterile 250 mL flasks containing 50 mL of either TAP media (control flasks) or nitrogen-depleted (N-TAP) media (replicate flasks). A single colony of strain cc124 is aseptically transferred into each flask under sterile conditions in a fume hood to prevent contamination. Both the control and replicant flasks were then monitored over 144 hours at set intervals (every 24 hours), during which multiple parameters were measured. During this period, parameters such as cell growth rate (cell density over each 24-hour interval), Pigment concentrations (chlorophyll A, chlorophyll B, and carotenoids per cell over each 24-hour interval), and the swim speed of the cell population were measured at an LCD period (at hour 24) and an HCD period (at hour 120).
Specification
To prepare TAP media suitable for Chlamydomonas reinhardtii, for 1-liter preparation, the media must for 1-liter preparation, the media must consists of 20 mL of 1 M Tris base (prepared by dissolving 30.14 g Trizma base in 250 mL DI water), 1.0 mL of phosphate buffer II (made from 10.8 g K₂HPO₄ and 5.6 g KH₂PO₄ in 100 mL), 10.0 mL of Solution A (containing 20 g NH₄Cl, 5 g MgSO₄·7H₂O, and 2.5 g CaCl₂·2H₂O in 500 mL), 1.1 mL of Hutner’s trace elements, and 1.0 mL of glacial acetic acid, with the remaining volume brought to 1 L using DI water. The final pH is adjusted to approximately 7.0 to support optimal algal growth. To prepare nitrogen-reduced TAP (N-TAP) media suitable for Chlamydomonas reinhardtii, for 1-liter preparation, the media must consists of 20 mL of 1 M Tris base (prepared by dissolving 30.14 g Trizma base in 250 mL DI water), 1.0 mL of phosphate buffer II (made from 10.8 g K₂HPO₄ and 5.6 g KH₂PO₄ in 100 mL), 10.0 mL of modified Solution A (containing 5 g NH₄Cl, 5 g MgSO₄·7H₂O, and 2.5 g CaCl₂·2H₂O in 500 mL), 1.1 mL of Hutner’s trace elements, and 1.0 mL of glacial acetic acid, with the remaining volume brought to 1 L using DI water. The final pH is adjusted to approximately 7.0 to support optimal algal growth.
Analysis
The growth rate (cell density) and Pigment concentrations (chlorophyll A, chlorophyll B, and carotenoid) per cell were quantified using a spectrometer, where samples were loaded into well plates and absorbance readings were taken at specific wavelengths (470 nm, 550 nm, 647 nm, and 663 nm). Blank readings from media alone were also collected to correct for background absorbance. The swim speed of the cell population was assessed by recording videos of live cells under a microscope at an LCD period (hour 24), then an HCD period (hour 120), and later analyzing them using the Fiji program (R-based tracking software). All collected data from the trials were organized in spreadsheets and analyzed using R code to calculate averages and interpret trends across conditions.
Future Works
Further studies into Chlamydomonas reinhardtii, will examine how its QS response to environment conditions, is affected by interchanging day-night cycle as they do have a natural circadian rhythm, which play a role in gene regulation.
Acknowledgement
Haire, T. C., Bell, C., Cutshaw, K., Swiger, B., Winkelmann, K., & Palmer, A. G. (2018). Robust microplate-based methods for culturing and in vivo phenotypic screening of Chlamydomonas reinhardtii. Frontiers in Plant Science, 9. https://doi.org/10.3389/fpls.2018.00235 Lee, D. Y., Park, J.-J., Barupal, D. K., & Fiehn, O. (2012). System response of metabolic networks in Chlamydomonas reinhardtii to total available ammonium. Molecular & Cellular Proteomics, 11(10), 973–988. https://doi.org/10.1074/mcp.m111.016733 Virtanen, O., Khorobrykh, S., & Tyystjärvi, E. (2020). Acclimation of chlamydomonas reinhardtii to extremely strong light. Photosynthesis Research, 147(1), 91–106. https://doi.org/10.1007/s11120-020-00802-2 Figure 1 created in https://BioRender.com
![]()

/prod01/fit-cdn-pxl/media/header-images/showcase-header.jpg)